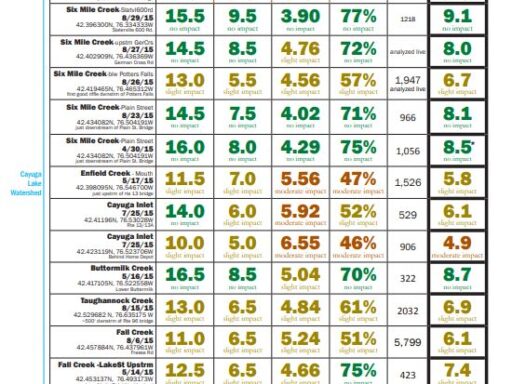
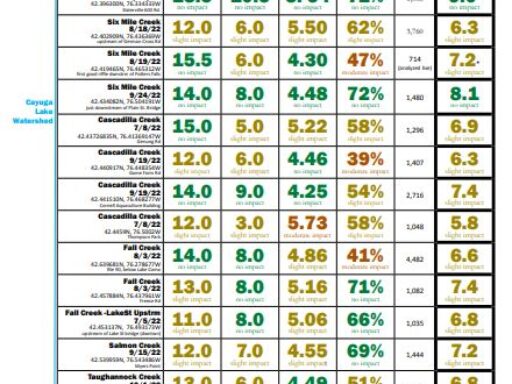
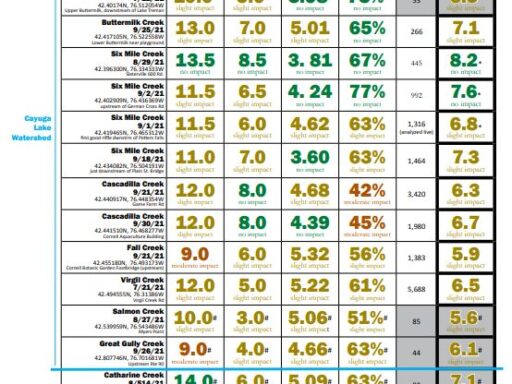
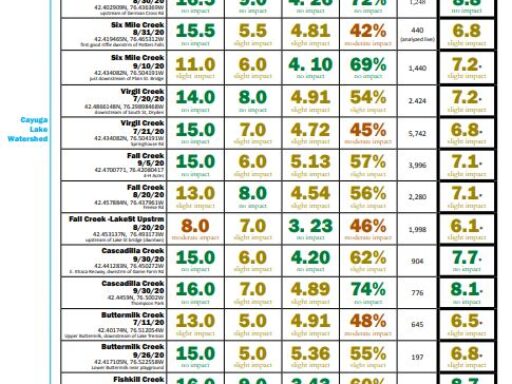
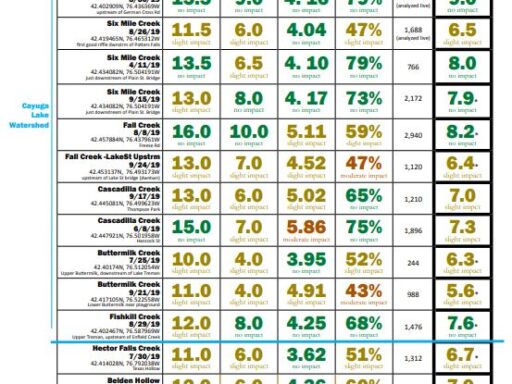
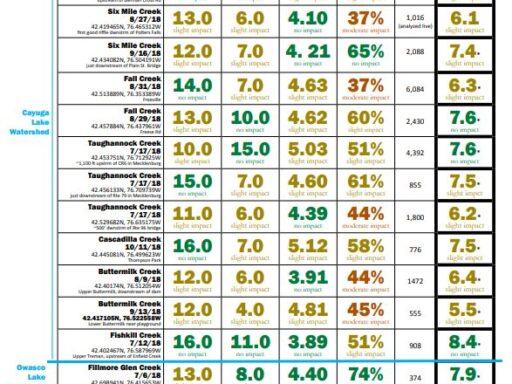
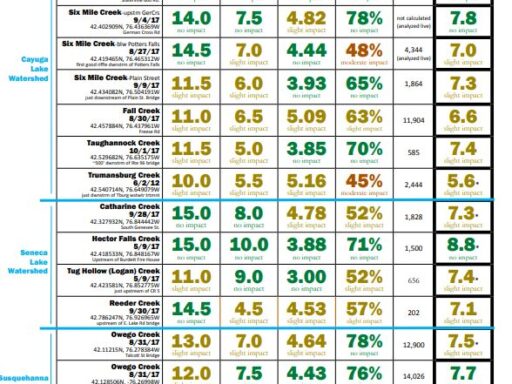
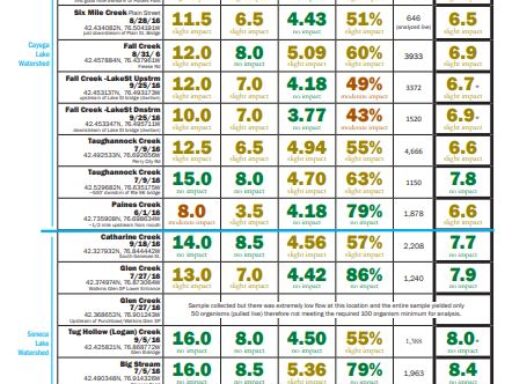
The metrics are explained followed by links to results (the most recent year for which results are completed on top). For more details about locations, or to see chemical monitoring data for the sites, visit the database.
Biological Monitoring Results
Biological Monitoring Results
Samples were collected and analyzed using a modified version of Hudson Basin River Watch Tier 3 Protocols.
Explanation of Metrics
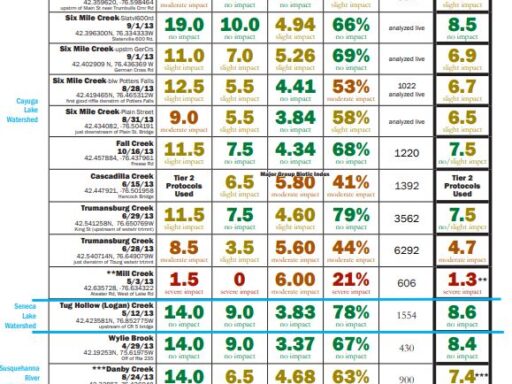
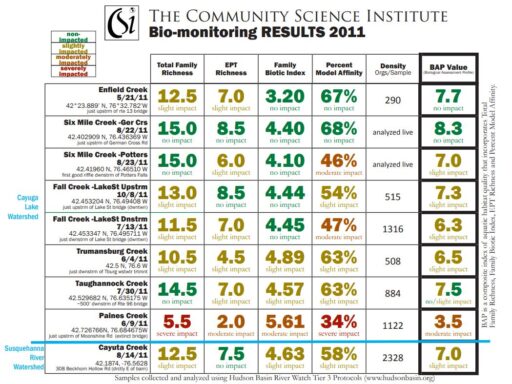
- Total Family Richness: A count of the number of different families of aquatic insects in the sample. It is an indication of how diverse the macroinvertebrate community is in the stream. The more different families of insects there are (higher richness), the healthier the stream.
Values: >13=non-impacted, 10-13=slightly impacted, 7-9=moderately impacted, <7=severely impacted - Family Biotic Index: Macroinvertebrate families vary widely with respect to how well they tolerate pollution. Some are very sensitive to pollution while others are quite tolerant of polluted conditions. This metric is based on the pollution tolerance of families of benthic macroinvertebrates on a scale of 0 to 10.
Values: 0-4.5=non-impacted, 4.51-5.50=slightly impacted, 5.51-7.00=moderately impacted, 7.01-10.00=severely impacted - EPT Richness: A count of the total number of macroinvertebrate families of mayflies (Ephemeroptera), stoneflies (Plecoptera), and caddisflies (Trichoptera) found in a sub-sample. These three orders of macroinvertebrates are considered to be mostly clean-water organisms, and the presence of several families from these three orders is generally correlated with good water quality.
Values: >7 = non-impacted, 3-7 = slightly impacted, 1-2 = moderately impacted, 0 = severely impacted - Percent Model Affinity: The New York State Department of Environmental Conservation has established what a typical, or model, community of benthic macroinvertebrates looks like in a healthy, non-impacted stream in New York. The Percent Model Affinity metric indicates how similar a BMI sample is to the model New York BMI community, as defined by the NYSDEC. It is based on the relative abundance of 7 major groups of benthic macroinvertebrates.
Values: >64%=non-impacted, 50-64%=slightly impacted, 35-49%=moderately impacted, <35%=severely impacted - Biological Assessment Profile (BAP): This metric combines Total Family Richness, Family Biotic Index, EPT Richness and Percent Model Affinity in a single metric called the Biological Assessment Profile (BAP). Each of these four metrics is converted to a value from 0 to 10 using a mathematical equation. The four converted values are averaged, and the average represents the overall water quality for the sampling location.
Values: 7.5-10 = non-impacted, 5-7.5 = slightly impacted,< 2.5-5 = moderately impacted, 0-2.5 = severely impacted
A Note About the Results:
In 2014 the Community Science Institute (CSI) continued to improve our volunteer bio-monitoring program by taking steps to make our BMI metrics more consistent with those generated by the New York State Department of Environmental Conservation. If you have been following this page closely, you may have noticed that some of the metrics and water quality values have changed, and the summary pages of results are labeled as “calibrated”. So what does this all mean? Read on to learn more.
CSI staff received family-level aquatic insect taxonomic certification from the Society for Freshwater Science.
Community Science Institute volunteers have been using Hudson Basin River Watch protocols to monitor water quality in streams by looking at benthic macroinvertebrate (BMI) communities since about 2002. In 2014, an opportunity arose for the current CSI bio-monitoring staff (Adrianna and Michi) to attend a 4-day training workshop on aquatic insect taxonomy with Watershed Assessment Associates (a professional taxonomy lab in Schenectady that is used by the New York Department of Environmental Conservation (NYSDEC) for analyzing some of their BMI samples). After the course, both Michi and Adrianna successfully pursued family-level aquatic insect taxonomic certification through the Society for Freshwater Science. This means that sample identification verified by Adrianna or Michi now holds a greater degree of credibility with agencies such as the NYSDEC.
We audited ourselves by having one replicate sample professionally analyzed and another analyzed by CSI bio-monitoring staff and volunteers. In doing this we discovered a flaw in Hudson Basin River Watch Protocols which led to a change in how we analyze volunteer data.
In an effort to further establish the credibility of our volunteer data, in 2014 we decided to pick one sample to send to Watershed Assessment Associates (WAA) to have professionally sub-sampled and identified to genus and species level. Another sample that was taken at the same time and location was evaluated by volunteers and CSI staff following Hudson Basin River Watch (HBRW) tier 3 protocols (half of it was analyzed live as preferred by some of our Six Mile Creek volunteers and half of it was preserved before analysis as per full HBRW tier 3 protocols). When we got the results back and compared our results with those of WAA, we were surprised to find that the resulting Biological Assessment Profiles (basically the average of a conversion of four different metrics to a scale of 1 to 10) for at least the preserved sample did not give an overall water quality rating that was comparable to the WAA results. The WAA results showed a “slight impact” (BAP=6.1) while the CSI results for the preserved sample showed a rating of “no impact” (BAP=7.7). On discussing this discrepancy with the folks at WAA, it turned out that the founder of that lab was also involved with Hudson Basin River Watch when they developed their protocols and he noted that the metrics used by both HBRW and the NYSDEC were developed for 100 organism sub-samples, even though HBRW in the end opted to establish the number of organisms to be looked at in a sample by using a State of Vermont-based protocol which calls for subsamples to be ¼ of the entire sample (i.e. usually quite a bit more than 100). He recommended that we take an electronic sub-sample of our data to back the numbers down to 100 organisms so that our results could more accurately be compared with theirs which were of a 100 organism sub-sample. On doing this, our BAP values for our preserved half of the CSI sample went from 7.7 to 6.7 and for the half of the sample that was analyzed live, it went from 7.1 to 6.6. After taking 100 organism electronic sub-samples, each of the three samples (our two and the one from WAA) gave a similar water quality rating of “slight impact.”
Through this effort to check our results against those of a professional taxonomy lab for the same site and date, we learned that in order to accurately use the metrics prescribed by the protocols that we use, we have to look at 100 organism sub-samples. That means that we could either figure out a way to randomly pull out exactly 100 organisms to sort and identify from a sample OR we could continue to follow the same protocols that we have been using and just take a randomized electronic sub-sample of the resulting data to back numbers down to 100. With the help of Ryan Dwyer (Laura, CSI chemist’s husband), we now have an Excel Spreadsheet that will do this for us. Continuing to follow the same protocols we’ve always used, which require looking at ¼ of the whole sample, we have the added benefit of recording a better picture of bio-diversity in our streams (than by only using a 100 organism subsample) as well as being able to get a sense of organism density. In very dense samples where one or two of the 12 squares of the sorting tray yield more than 500 organisms, volunteers might consider stopping at one or two squares (or 1/12 or 1/6 of the entire sample) since, according to discussions with AJ Smith of the NYSDEC, at that point one should have a pretty good sense of what different taxonomic groups are present in the sample. We also learned from AJ that higher density of organisms is often linked to nutrient loading so we will now be reporting density values (per sample) on our results posters.